Welcome to the Vizome documentation
URL: https://vizome.readthedocs.io/
Source: https://github.com/ohsu-heme/vizome_docs

Vizome is an interactive, context-aware knowledge discovery platform created by Libbey White, Beth Wilmot, Dan Bottomly, and Shannon McWeeney at Oregon Health & Science University.
There are three datasets currently available in Vizome:
- Beat AML (Tyner et al., Nature 2018)
- Crenolanib (Zhang et al., Nat Commun. 2019)
- Chronic neutrophilic leukemia (Zhang et al., Blood 2019)
The links in this documentation point to the Beat AML Vizome, but the information is similar for the other two datasets.
Much of this documentation can also be accessed in Vizome by clicking the  button on each page.
button on each page.
Summary of Modules in Vizome
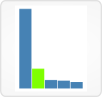 1. Sample attributes view
1. Sample attributes view
This view provides several methods to define a global filter that will limit browsing in Vizome to a subset of samples.
 2. Variant filters and search
2. Variant filters and search
This page presents options for defining global filters based on variant properties, as well as options for searching for variants.
 3. Gene sets
3. Gene sets
This view allows you to define a set of genes, or to select a pre-defined set, which you may then explore in a variety of ways.
 4. Mosaic view
4. Mosaic view
This view displays data related to samples, including clinical and variant, and fusion data (if applicable).
 5. Individual view
5. Individual view
This view displays variants and inhibitor results for an individual, one patient at a time.
 6. Compare gene variants view
6. Compare gene variants view
This view allows you to compare a set of genes selected via DNA variants found in either individual samples or groups of samples.
 7. Expression stratification view
7. Expression stratification view
This view displays RNA-Seq expression for a given gene and various sample attributes.
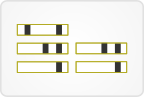 8. Chronology view
8. Chronology view
This view displays DNA variants for a given gene, sorted by sample date for each patient.
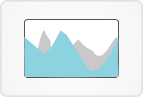 9. Gene model view
9. Gene model view
This view displays gene models, variants, DNA coverage, miRNA, CTCF, target regions, DNase, H3K27ac, and, if applicable, RNA coverage, fusions, splicings, and a heatmap of RNA-Seq expression.
 10. Protein view
10. Protein view
For a given gene, this view displays Pfam domains of the protein it codes, and variants in the study population.
 11. Inhibitor view
11. Inhibitor view
Results from inhibitor testing, variant data, and normalized RNA-Seq expression data appear in this view.
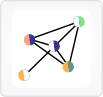 12. HitWalker
12. HitWalker
This view provides an interface to the HitWalker program, which ranks genes containing variants with respect to functional data or supplied gene sets.
Getting Started
See tutorials page for instructions and examples.